In my two previous posts, I’ve been sharing some tidbits that I learned at the PRB Policy Communication Workshop. In my first post, I aimed to motivate you to think about the broader impacts of research, especially considering the unique role researchers play within the process of policy formation or change. In my second post, I discussed three different outlets–aside from academic journals–where researchers can share their findings with the public. This week, in my third and final post about policy communication, I will share some tips that I learned about conversational writing. Special thanks to Craig Storti for his enlightening presentation about some bad habits that I picked up in grad school!
Disclaimer: This blog post contains several cat puns. This may result in audible groaning and face-palming. Reader discretion is advised.

Academic Jargon and Dense Prose
It may seem obvious that we should avoid academic jargon when writing for non-technical audiences. As I said previously, abstract concepts such as macro- and micro-level processes or statistical methods are not well understood outside a specific discipline. We are also often told that we should stop using words such as ‘utilize’ when we could easily substitute ‘use.’ But even if we are acutely aware of these bad habits, here are two other occupational hazards that I did not consider before the workshop: 1) Nominalization and 2) Noun Compounds:
Nominalization is when we transform a verb into a noun. For example, nominalization itself is a noun that was derived from a verb–i.e., ‘nominalize.’ Another example is the word ‘investigation’, which is from ‘investigate.’ Sentences that contain nominalized verbs can be weaker and less concise than sentences that use the actual verb.
A Noun Compound is when we use a consecutive string of two or more nouns in a sentence. For example, ‘Policy Communication Workshop Fellowship’ or ‘national community health operations research technical working group.’ Excessive use of noun compounds can result in dense writing that is difficult to understand.
Example
To demonstrate how easy it can be to both nominalize our verbs and string several nouns together, I wrote a hypothetical introduction to the cat meme inequality study<–noun compound!–that I used as an example in my previous post. Nominalizations are underlined; noun compounds are in red (excluding the phrase ‘cat meme’ alone); and jargon is in blue. Puns are italicized 🙂 :
Differences in purr household consumption of cat memes have been dramatically increasing over the past half-century, and research suggests that this growing disparity is due to incongrooment access to cat memes. Informed by this body of research, my study utilized data from the Cat Meme Survey of Households and Families and found that legislative pawlicies have, in part, catapulted these cat meme inequality access issues. Right meow, cat meme pawlicies are littered with supurrrfluous loopholes fur the rich and privileged. However, my research indicates that these catastrophic inequalities in cat meme access can be mitigated if pawlicymakers consider the implementation of laws or clawses that focus on the inadequacy of cat meme access fur more disadvantaged households through the creation of cat meme inclusion zones, which would allow fur the dissemination of more provisions fur those who are in need.
Tips for avoiding dense prose
The simplest way to avoid nominalizations is by restoring the verb. For instance, the first sentence of my example could be changed to “Rich households consume more cat memes than poor households…” Alternatively, the sentence could be changed to “Households are consuming cat memes at a different rate…” The latter example uses the gerund form of the verb.
The benefit of correcting nominalizations is that you will likely break up noun compounds, like I did in my first example:
Original: Differences in purr household consumption of cat memes have been dramatically increasing over the past half-century…
Corrected: Rich households consume more cat memes than poor households, which is a trend that has been increasing over the past half-century.
Another way to fix noun compounds is by including a preposition such as ‘of’, ‘in’, ‘to’, and ‘for’:
Original: However, my research indicates that these catastrophic inequalities in cat meme access can be mitigated if pawlicymakers consider the implementation of laws or clawses that focus on the inadequacy of cat meme access fur more disadvantaged households through the creation of cat meme inclusion zones, which would allow fur the dissemination of more provisions fur those who are in need.
Corrected: My research indicates that access to cat memes across households is inadequate. Policymakers should consider implementing laws that help more disadvantaged households gain access to cat memes. For example, by creating incentives to encourage builders and investors to provide more households with equal access to cat memes, or restricting builders and investors from accessing permits unless they agree to these terms, which is often referred to as “inclusionary zoning.”
It gets better with practice
I was surprised by how difficult it was to correct nominalizations and (especially) noun compounds at the workshop. I found that some of my resistance to removing noun compounds is that it can result in longer sentences. But unless I am writing for an academic journal, the value of writing more concisely is lost when my audience does not understand what I am writing about. It’s a skill that I will have to continue to practice and be more thoughtful about in the future. I encourage you to do the same!



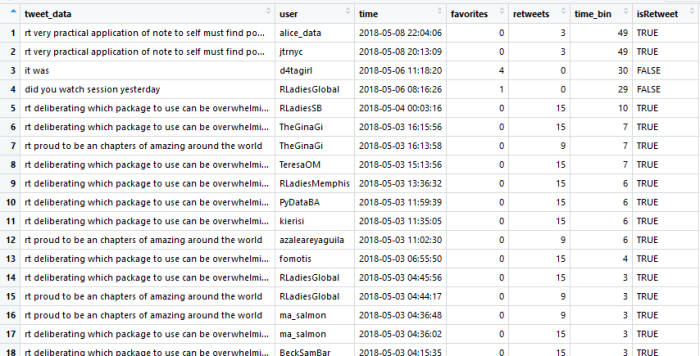
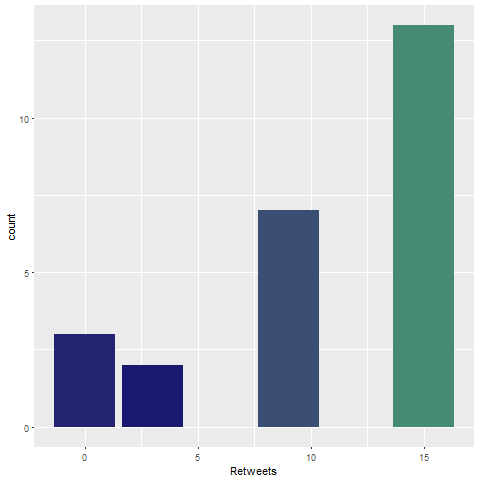


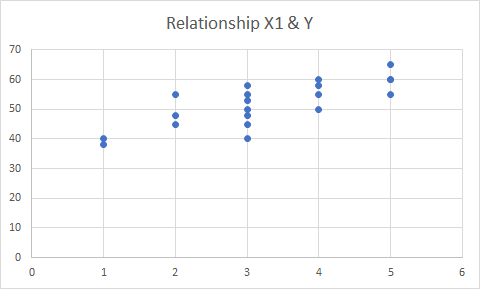
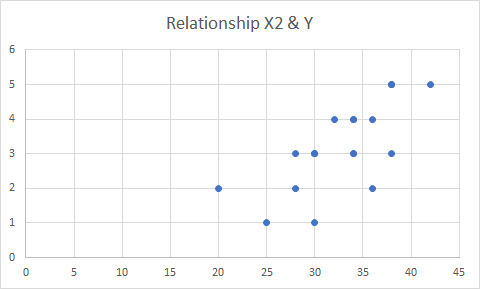




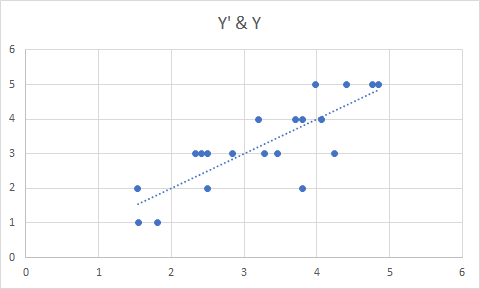
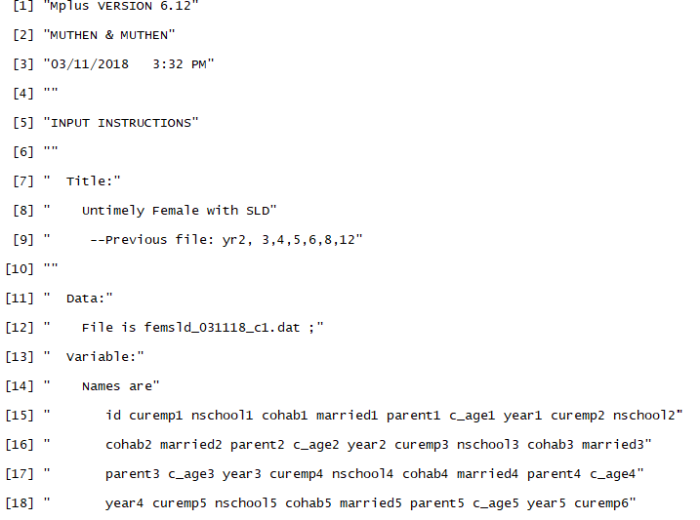
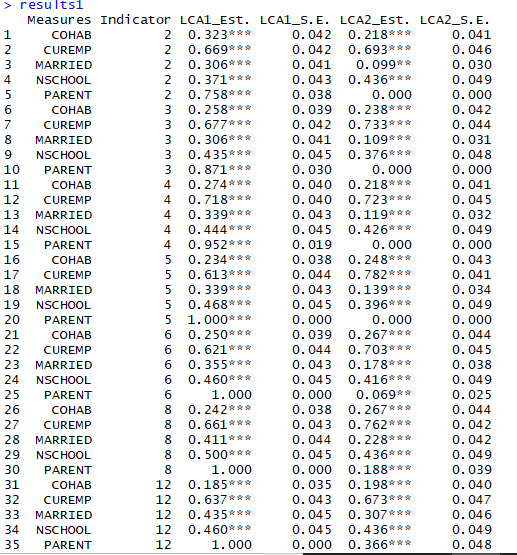
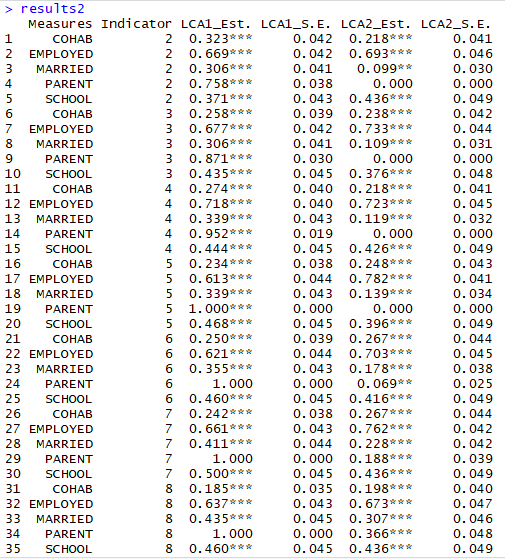
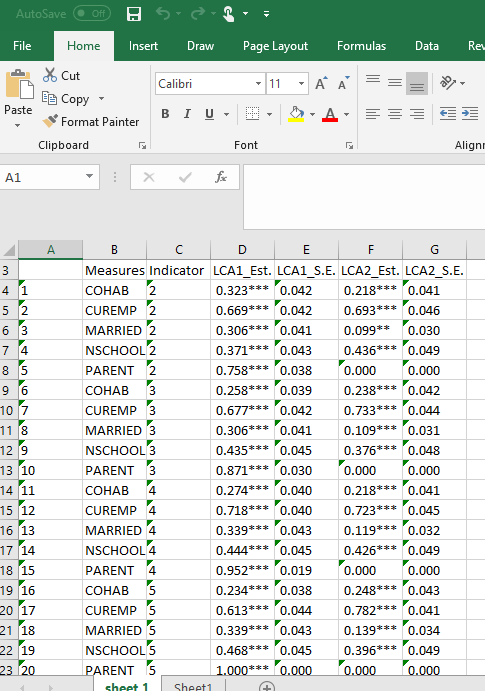
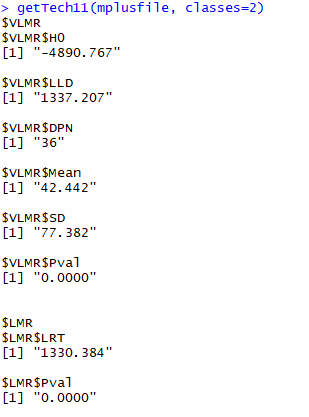
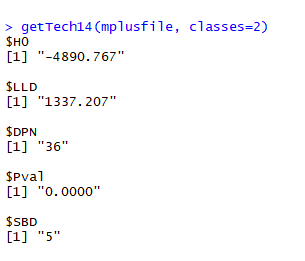
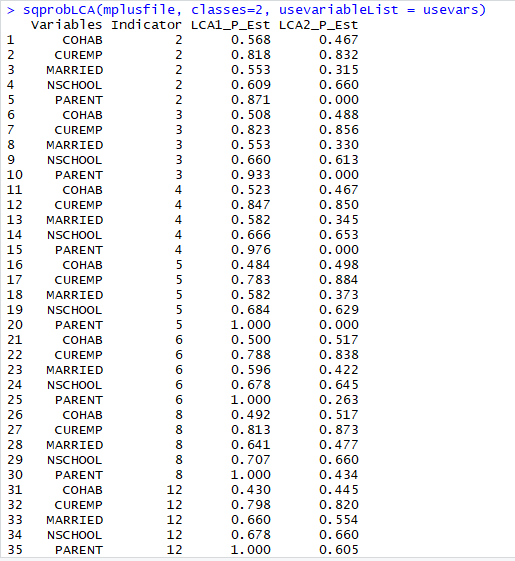


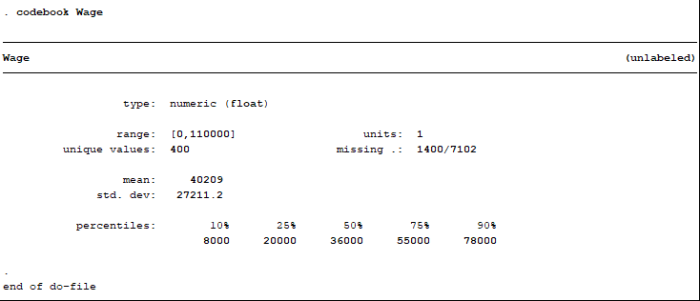
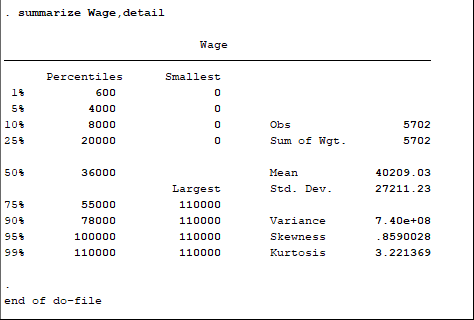
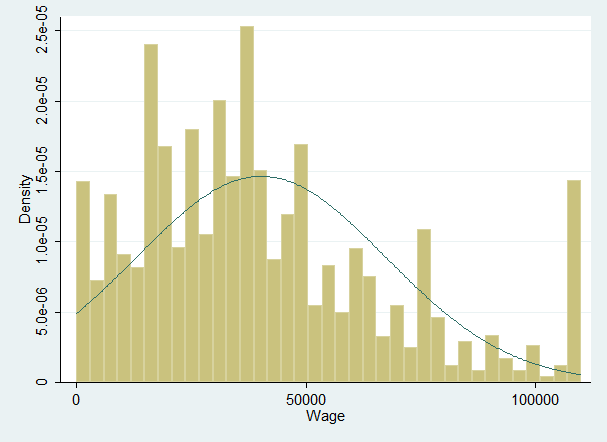
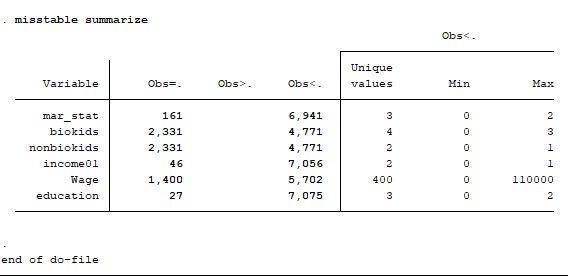
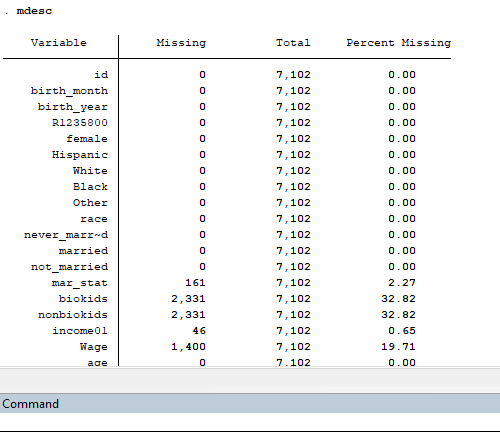
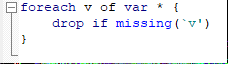
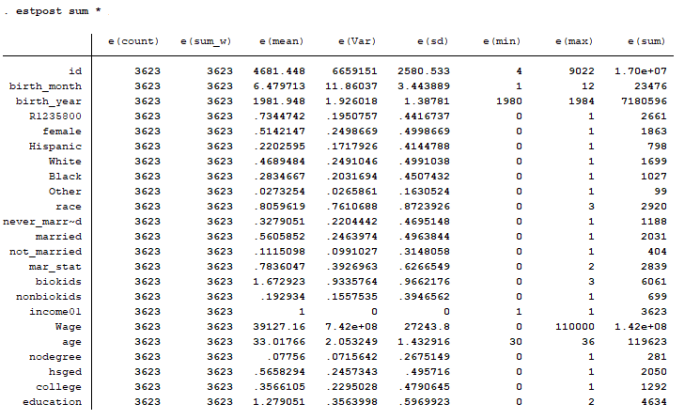
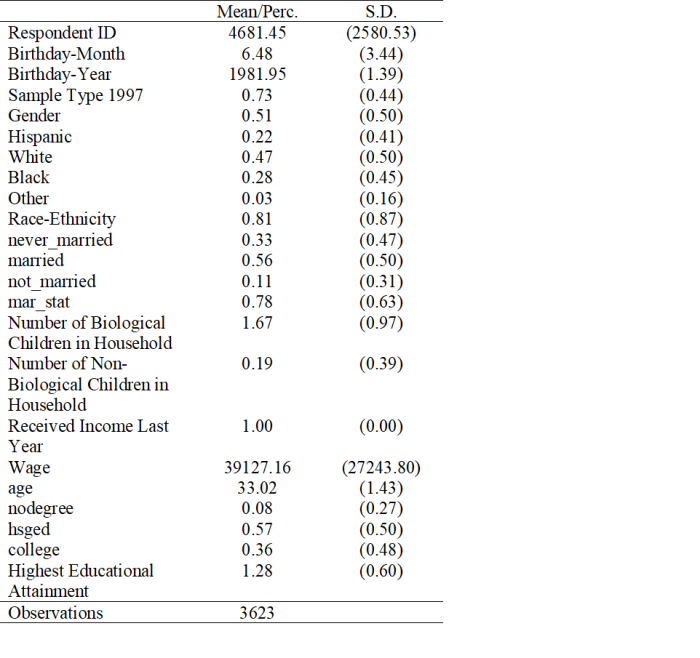 Note that not all of the variables are labeled. You will want to do this before you submit an assignment or use the table in a research paper.
Note that not all of the variables are labeled. You will want to do this before you submit an assignment or use the table in a research paper.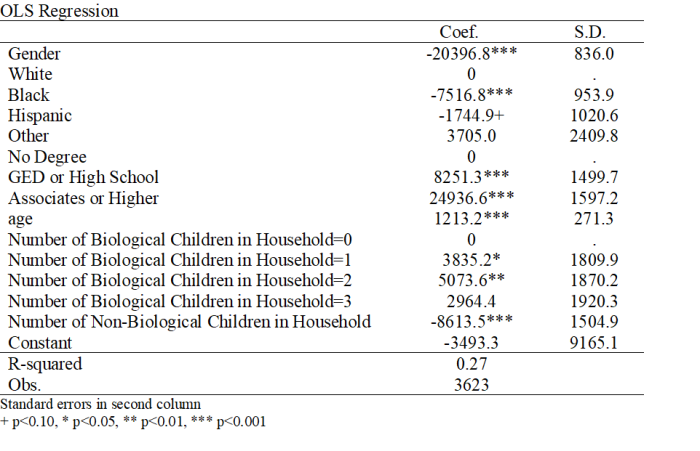 I show you how to do this toward the end of the program, after I go through quick explanations of how to check basic OLS assumptions.
I show you how to do this toward the end of the program, after I go through quick explanations of how to check basic OLS assumptions.